A few months back I worked out how to do a bacterial transformation, and then I (successfully) preformed the procedure. I did this at BOSSLAB, which is in Sommerville, MA. If you are in the area, check it out! This procedure is memorialized in the BOSSLAB lab notebook, which I will scan and upload relevant portions of to the downloads page, but one of the goals of BOSSLAB (??? at best) seems to be to communicate what we do there to the world. This is my writeup on what I did and how I did it.
What is Transformation?
Transformation is the process of adding genetic material to a bacteria. The phrase comes from an experiment by Fredrick Griffith in 1928, where he discovered that non-pathogenic strains of Strep. pneumoniae could be “transformed” into pathogenic strains. This was done by adding dead (heat-killed) pathogenic Strep. pneumoniae to the non-pathogenic strains. When this mix was injected into mice, the mice died. Later, the Avery-MacLeod-McCarty experiment showed that DNA caused this transformation.
For bacteria, the most convenient way to add genetic material is with plasmid DNA, which is a little circle-shaped piece of DNA. There are a few really important things to know about choosing a plasmid, most of which I (fortunately) got to gloss over.
As you can see on the right hand diagram, there are three major parts to a plasmid. An origin of replication, which is dependent on the target organism (ori-> on the left diagram). This is where the cell begins the replication of the plasmid, if it is going to make more. Another important factor in plasmid selection is some kind of selection marker (ampr on the left diagram). Both of these plasmids use an antibiotic resistance marker. This is important so that when you have all the transformed and not transformed bacteria mixed up, you can selectively kill all the ones that do not have the antibiotic resistance gene. The final consideration are the regions that can be cut by restriction enzymes, which are also called restriction enzyme digest sites. These are used to cut the plasmid open, so you can add genes to an “empty” plasmid vector. When you have a lot of them together, near each other, it is generally called a polylinker or multiple cloning site. On the left hand diagram, if you “digested” (cut) the plasmid with EcoR1, you would have two EcoR1 compatible ends on the plasmid. You could then add a gene with EcoR1 compatible ends.
For me, all I did was buy the plasmid on the left. The goal was to successfully complete a transformation, not do ligate a gene into a plasmid. That might be the next step, but trying to do it as an initial foray into genetic engineering or running my own protocol completely independently would add too many uncertainties to the protocol and make it impossible to debug with the materials I had. For you embedded folks, this would be like trying to hand code assembly with no debugger and no verification of the program memory.
How do you do it?
Bacterial transformation is tricky. You can put plasmid in a tube with some bacteria and swish it around all day, and probably nothing will happen. Thats because the outside of bacteria is a membrane which keeps things that should be out, out, and things that should be in, in. Basically it’s like their skin. And the plasmid needs to go through it. To do this, we have to break up the skin so tiny pores open. When bacteria are in this state, they are called competent.
There are various recipes for making chemically competent cells. I used CaCl salts, but I hear tell that TSS (Transformation Storage Solution paper by Chung et al. here) is more efficient. Unfortunately, I had no success with TSS because of (probably) a snafu with a thermocouple, and I didn’t have time or money to do it again (see what went wrong). Both of these solutions have the same effect; the positive ions in the solution neutralize the (negative) charge on the membrane, and the negative charge on the outside of the DNA.
Once the cells are competent, and chilled, DNA is added. Later, they are “heat shocked”, which (possibly?) expands the holes in the membrane and allows the DNA to enter the cell. Afterwards, they are allowed to recuperate, and then a tiny amount of bacteria are spread onto a plate with the antibiotic. The goal here is that the bacteria without the plasmid will die, leaving you colonies of the transformed bacteria (transformants).
Results (and what went wrong)
First, a picture!
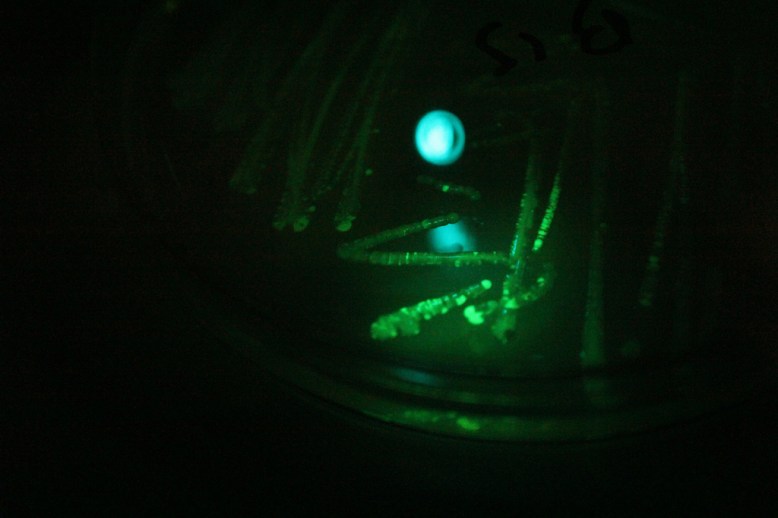
Transformants under blue light with a green filter. Green light is produced by the fluorescence, and then only green light is let through the filter.
Although I got the procedure to work, I had less than stellar results. I attempted transformation twice. The first round had five trials, all of which completely failed. Some of these used TSS and some of these were CaCl. The reason these failed is probably because the water bath was not hot enough. Since we didn’t have a temperature controlled water bath, we used a large beaker of water on a hot plate instead. The idea is that we would heat it up to about the right temperature (42C) and then turn it off. since water retains heat pretty well, we thought it would stay near enough to 42C to be used for the transformation. HOWEVER, on my second attempt at transformation (only two trials this time!) I was measuring the temperature of the water with the same thermocouple we had used before…and I discovered that it was VERY wrong. I believe the thermometer was on the low side, reading -20C in ice water, so the water for the heat shock was probably around 60C-way too hot. When I discovered this in the second trial, I had to very quickly figure out a way to get some 42C water! My bacteria were incubating and I was eager to get on with the heat shock.
I knew from making antibiotic plates that at about 50C, it is safe to add antibiotic and that the media will be hot, but bearable to hold. So I heated some water until it was uncomfortable to touch, and then cooled it down some. Estimating that my body temperature was 35, I knew the water would feel very warm, but not too warm to touch. Using this, I guesstimated that I had a good temperature and preformed the heat shock.
All of this can be avoided by buying a (literally) $3 thermometer from the dollar store.
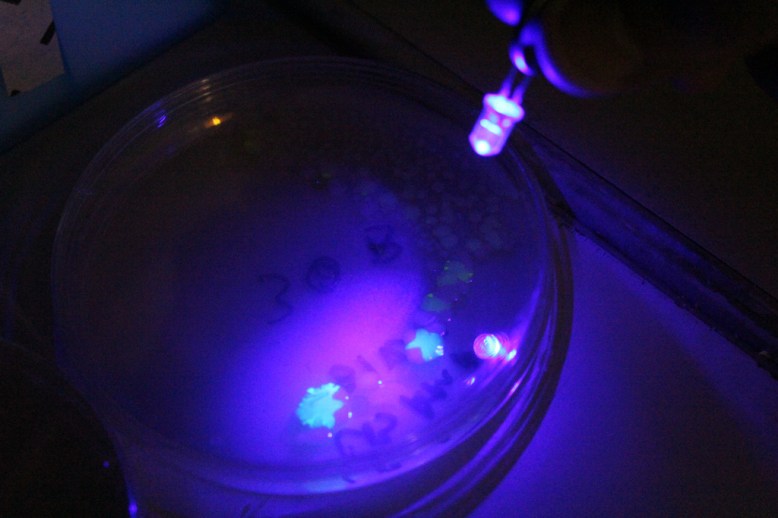
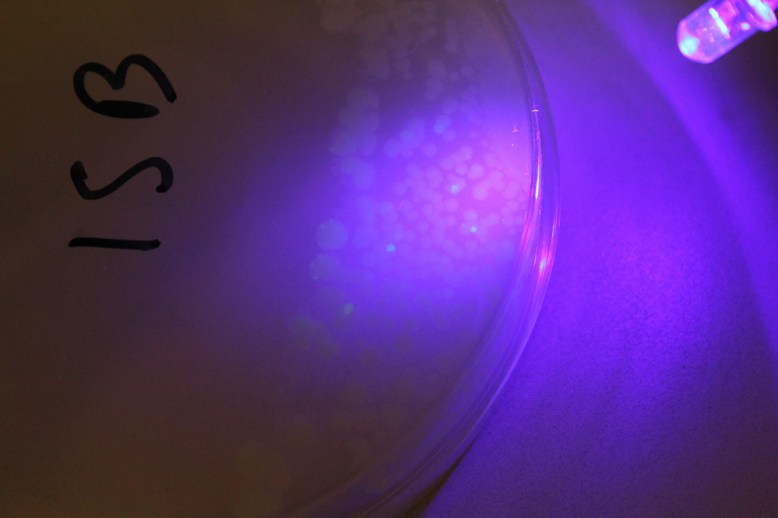
Another important step that I forgot to figure out before I began was how to verify the GFP was being produced. It turns out that this variant of GFP fluoresces under blue light. Since there were batteries and a few blue LEDs handy, I used those.
The other, sneakier problem, was the lack of antibiotic efficiency. This caused satellite colonies (or maybe just amp-R colonies) to proliferate over the plate. I have a few theories about why this could have happened:
- There is some sort of ampR contaminant that got moved from plate to plate
- The antibiotic was bad. I tested it before the first attempt, but it could have gone bad in the mean time (sketchy fridge is sketchy), or it could have been damaged in pouring. I didn’t have enough plates to test if the plates I poured worked, and to do all the transformations.
- The bacteria incubated too long, and they destroyed enough ampicilin locally to allow satellite colonies to form.
I think a combination of two and three are most likely, particularly two. In the first set of trials I also saw slow growth, but no GFP. This would indicate that there is something wrong with the ampicilin, possibly that it was not concentrated enough, or that it was damaged. I think three is also likely because I mostly saw non-GFP growth near the GFP bacteria, or where I pipetted the transformation solution onto.
Procedure (specifically, how do you do it)
The actual procedure I followed, can be found at this link to a pirate pad where I planned the transformation out this summer. I will also put a nice PDF version (including some of this reference material) on the downloads page.


Hey! Great to see that you got it working in the end. Transformation is an exciting milestone in self-taught SynBio. 🙂
When it comes to “best method”, I highly recommend TSS, precisely because you *do not need to heat-shock*. You also do *not* need to add DMSO if you can’t get it. And because the only other ingredients are PEG-3350 (a common laxative; brands such as GoLytely and MiraLax are pretty pure PEG3350) and MgSO4 (Epsom Salts), it’s really easy to do after a simple shopping trip in town.
So, next time you might consider TSS because it’s faster, easier and more reliable. No water baths, and you can make it as simple as “dilute overnight cells in 2x TSS, add DNA, walk away for an hour, select on antibiotic plates” if you’re willing to accept poor efficiency. 🙂
Thats actually my next project! I have DSMO, although now that you mention it I will have to try some without it. Thanks for the tip!
Great site! I am loving it!! Will be back later to read some more. I am taking your feeds also